Thesis
Monoclonal antibodies are one of the fastest-growing segments of the pharmaceutical industry, accounting for 25% of new drug approvals in 2025. The pace of regulatory adoption of monoclonal antibodies accelerated through 2024, in which 21 antibody therapeutics were granted a first approval in one country or region. The drugs also entered mainstream recognition among pharmaceuticals when monoclonal antibodies were used for the treatment of COVID-19 in some outpatient cases.
Antibodies are Y-shaped glycoproteins that are produced by the human immune system. Several structural properties differentiate them from traditional small-molecule drugs, establishing their advantages as a therapeutic modality. Antibodies have a much higher specificity relative to small molecules because there is more engineerable surface area on the antibody. This is especially important because safety issues from non-specific binding account for 28% of small molecule drug failures. Further, antibodies have a serum half-life in the bloodstream of 10 to 21 days, while small molecules only have a half-life of a few hours. Together, these properties enable less frequent dosing with improved patient outcomes.
However, the process of discovering new antibody drugs remains inefficient and relies on a combination of two approaches. The first, called rational design, describes researchers relying on preexisting knowledge of antibody structure to design targets for antibody binding. The second, directed evolution, refers to researchers starting with an antibody and randomly mutating the gene that encodes for it to then screen the variants for desired properties.
A computational alternative to these methods is becoming viable as three independent technology trends converge. As of February 2026, AI training costs have declined at approximately 10x per year, with the cost per floating point operation per second (FLOPS) in 2025 decreasing approximately 74% since 2019. These improvements enable the training of large generative models on protein structure and sequence data. Second, DNA synthesis costs have fallen from $25 per base in 1999 to around $0.3 per base in 2025, meaning computationally designed antibodies can be synthesized and experimentally validated at costs that enable high-throughput iteration instead of trial and error. Finally, the release of AlphaFold2 in July 2021 proved the concept of atomic-level protein structure prediction for the first time. Subsequent model iterations built on top of this technology demonstrated that not only is protein structure predictable, but it can also be designed.
Chai Discovery leverages this progress to build generative and predictive models that learn design principles for antibody engineering. Its flagship model, Chai-2, allows users to provide an antigen structure and have the model provide antibodies that bind to that antigen. This generative antibody design process compresses the antibody discovery cycle from 12 to 24 months to four to eight weeks.
Founding Story
Chai Discovery was founded in March 2024 by Joshua Meier (CEO), Jack Dent (President), Matt McPartlon (CTO), and Jacques Boutreau.

Source: TechCrunch
Coming from a family of physicians, Meier always had an interest in science. At 16, he was a finalist in the 2012 Google Science Fair, looking at artificially generated stem cells. In high school, he founded Provita Pharmaceuticals, a company focused on genetically engineering mosquitoes to deliver a vaccine for the West Nile Virus. During his time as an undergraduate at Harvard, Meier trained at the Feng Zhang lab, where he worked on the lab's genome-wide CRISPR screening efforts.
After graduating from Harvard, Meier was recruited to OpenAI, where he was exposed to advances in natural language processing and became interested in applying these models to understand DNA, “the language of biology.” He went on to work at Meta in its generative biology group, where he helped develop ESM1, the first transformer protein-language model. He then spent three years at Absci, supporting its AI efforts, eventually becoming the Chief AI Officer in 2023.
Before founding Chai Discovery, Meier and Dent were classmates at Harvard, where they studied computer science together. After graduating, Dent went on to work at Stripe, where he helped launch Stripe Link and Stripe Capital. However, they would meet every three to six months, with Meier updating Dent on his research progress.
McPartlon worked with Meier at Absci as part of its AI research team, working on de novo antibody design modeling. Before Chai Discovery, he was recruited to Proxima (VantAI) to work on protein-protein interaction models. Boutreau studied at Jerome Waldispuhl’s RNA bioinformatics research group at McGill University, working on the molecular affinity optimization using generative models. After graduating, he was recruited as an AI scientist at Aqemia, where he worked on small molecule design.
The startup began amid conversations between its co-founders and Sam Altman. Altman wanted to recruit Meier to work on an OpenAI proteomics spinout. However, Meier felt that proteomics technology “was not quite there yet.” In 2024, Meier and Dent contacted Altman to pick up the conversation about working on the proteomics company that they had discussed with Altman. This conversation turned into Chai Discovery, where OpenAI became one of the first seed investors.
Meier made the bet that advances in language models would become useful for drug discovery. The vision for the company was to make biology less like experiment-based science and more like engineering. In Meier’s view, part of the approach’s advantage was avoiding tight lab integration that was common in the previous generation of AI biotech companies, and instead becoming a “portable AI platform.” The team went on to recruit leading AI research scientists to develop new tools for the industry. In September 2024, Chai Discovery revealed Chai-1, an open-source foundation model for molecular structure prediction that performs similarly to AlphaFold.
After the launch of Chai-1, adoption grew as researchers began integrating the model in research workflows. In August 2025, Chai Discovery announced a $70 million Series A and unveiled Chai-2, a model for de novo antibody design with a near 20% hit rate. As part of this round, Mikael Dolsten, Pfizer’s former Chief Scientific Officer, joined the Board of Directors of Chai Discovery.
Product
The core product philosophy of Chai Discovery is a “computer-aided design suite” for molecules. This concept is borrowed from mechanical engineering, where tools like AutoCAD transformed the process of generating technical drawings and specifications. Chai Discovery’s stated mission is to enable a similar transformation in drug discovery: transforming biology from random trial and error to an engineering discipline.
Chai-1
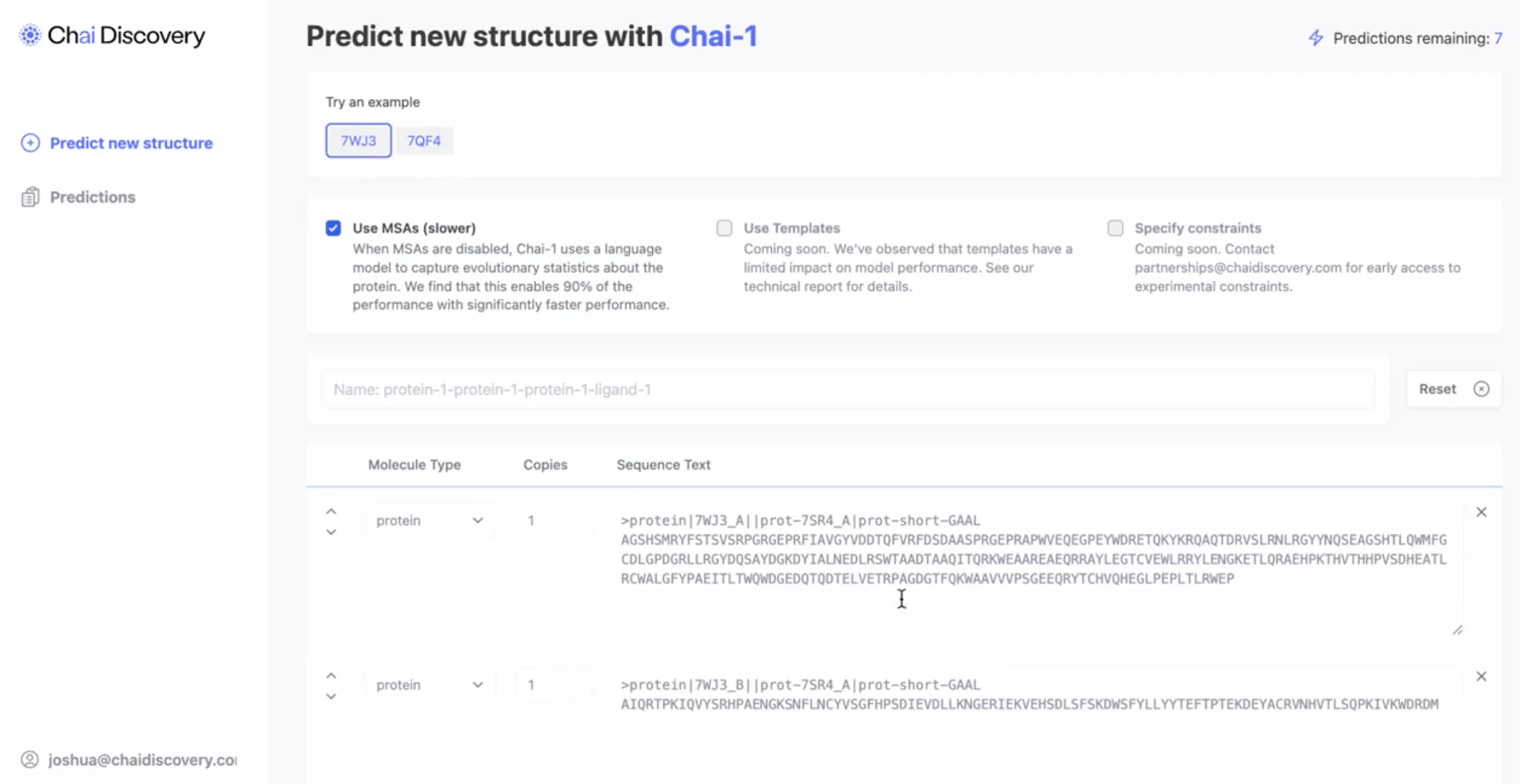
Source: Chai Discovery via X
Chai-1 is a multimodal foundational model designed to predict the structure and interactions of biochemical molecules. Released as open source, the model supports structure prediction across proteins, small molecules, DNA, RNA, and covalent modifications. This is a critical assessment in monoclonal antibody drug discovery, as protein-ligand interactions form the basis of the mechanism of action for the treatments.
Meier describes this product as an “atomic-level microscope” used to predict atomic interactions when rationally designing drug targets. Chai-1 was found to perform at or above state-of-the-art models across industry-standard biomolecular structure prediction benchmarks.
When tested on 152 antibody-antigen complexes held outside the training set, Chai-1 achieved DockQ scores above 0.8 (good binding interface prediction) on 17% of the structures. This is approximately 2x better than AlphaFold 2's performance on the same benchmark. DockQ scores quantify predicted versus actual binding interface accuracy on a 0 to 1 scale. A score above 0.23 is considered acceptable, while a score above 0.8 is considered highly accurate.
Chai-2
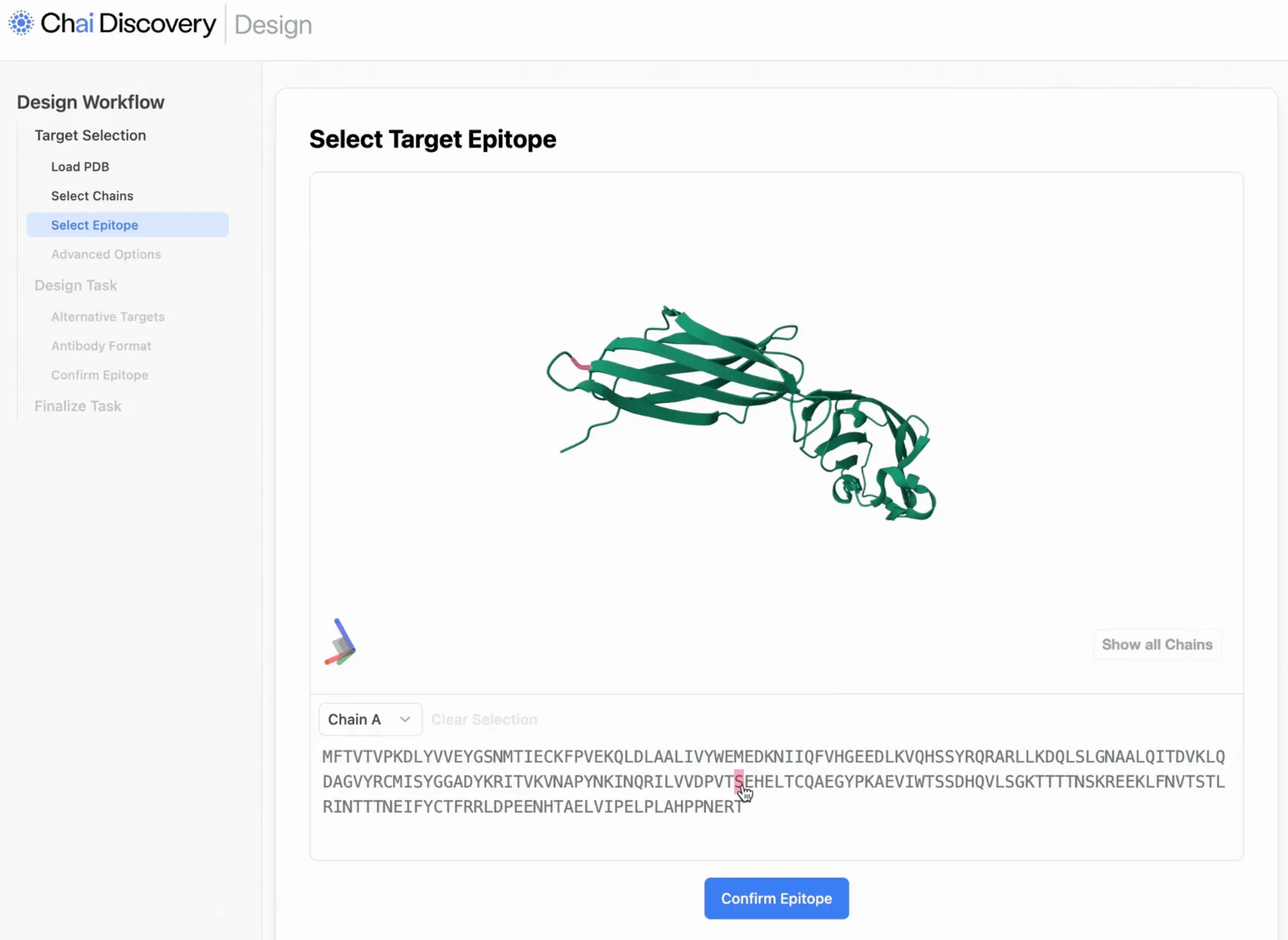
Source: Chai Discovery via X
Chai-2 is an end-to-end model composed of multiple submodels focused on de novo antibody design. Meier describes the Chai-2 product as “Photoshop” for antibody design. This involves generating new antibody sequences from scratch rather than optimizing existing antibodies.
Chai-2d (design) is the submodel used for de novo antibody sequence generation against arbitrary and pre-specified protein targets. The design task allows the user to specify the epitope (region to bind) and modality format (full-length IgG, VHH, bispecific). The model uses an all-atom generative protein model framework, which allows it to design the amino acid backbone and side-chain atomic structures.
Chai-2f (folding) is an enhanced structure prediction model that has a similar architecture to Chai-1. Chai-2f is trained on synthetically predicted structures and the protein structures in the PDB. When benchmarked on the same 152 antibody-antigen complexes used to evaluate Chai-1, Chai-2f achieved DockQ scores above 0.8 on 34% of structures, which is 2x the performance of Chai-1.
The integrated architecture creates an antibody engineering workflow. Chai-2d generates antibody sequences that target a specified epitope, and then Chai-2f predicts the resulting antibody-antigen complex structure. These designs can then be iteratively refined based on the predicted binding interfaces.
According to announcements made by Chai Discovery, this approach achieved approximately 20% experimental hit rates across 52 diverse targets. This represents a 100x improvement over previous computational antibody design methods, which have success rates around 0.1%.
Market
Customer
Chai Discovery’s main customer segment is large pharmaceutical and biotechnology companies developing antibody therapeutics. In January 2026, Eli Lilly announced a partnership with Chai Discovery to deploy the company’s models across its therapeutic portfolio. As part of this licensing agreement, Eli Lilly pays an annual access fee in the mid-eight figures.
Beyond the larger pharmaceutical companies, Chai Discovery also sells its models to government and academic research institutions, and emerging biotech companies. Chai Discovery announced that it would participate as part of the UK government's OpenBind initiative alongside Isomorphic Labs and Genentech. Many research papers also report using Chai-1 in their methodologies. In 2025, 304 papers cited Chai-1, demonstrating academic research use. In addition, Meier reports that Chai Discovery received “hundreds of early access requests” when Chai Discovery announced Chai-2. This includes biotechnology startups developing therapeutics.
Market Size
The antibody market is growing in line with the demand for targeted therapies. The global market for monoclonal antibody therapeutics reached approximately $288 billion in 2024 and is projected to grow to $628 billion by 2035. This market has grown at a CAGR of 8-10% from 2015 to 2025, driven by expanding indications in oncology, immunology, and rare diseases.
The market has seen many breakthroughs since the introduction of the first method of generating monoclonal antibodies by Nobel Prize winners Georges Köhler and Cesar Milstein in 1975. In 2025, over 150 monoclonal antibodies have received FDA approval since the first therapeutic antibody (Orthoclone OKT3) in 1986. In 2025, 23% of newly approved drugs were antibody therapies. The market continues to grow as new antibody modalities are introduced, such as the following:
Single-Chain Variable Fragment Antibodies (scFv) are being developed for their higher efficacy and ease of manufacturing. For example, brolucizumab, developed by Novartis, is a humanized scFv used for the treatment of age-related macular degeneration.
Nanobodies are being introduced because of their improved efficacy, significantly higher stability (stable at room temperature), and ease of manufacturing. Caplacizumab, developed by Ablynx, was the first nanobody approved by the FDA in 2019. The company was acquired by Sanofi for €3.9 billion in 2018 for its nanobody platform.
Antibody mimetics are small, non-antibody organic compounds engineered to bind specific antigens with high affinity. Antibody mimetics have similar advantages to scFv and nanobodies, with the added benefit of being extremely flexible.
As these next-generation antibody platforms mature and become clinically validated, the market for antibody engineering tools like those provided by Chai Discovery to develop these antibodies will expand with it.
Competition
Generative Models
Boltz: Boltz is a family of open-source models for biomolecular interaction prediction. In November 2024, Boltz-1 was announced as the first fully open-source model to approach AlphaFold3 accuracy. In June 2025, Boltz-2 was announced as the first deep learning model to approach the accuracy of physics-based free-energy perturbation (FEP) methods. In January 2026, Boltz announced a $28 million seed round at an undisclosed valuation and its launch as a public benefit corporation (PBC). As part of this round, it announced a multi-year partnership with Pfizer to build exclusive models for structure prediction, small-molecule affinity, and biologics design.
EvolutionaryScale: EvolutionaryScale was founded by Alexander Rives, who led Meta’s AI protein team, in 2023. EvolutionaryScale raised a $142 million seed round in June 2024 at an undisclosed valuation, led by Nat Friedman, Daniel Gross, and Lux Capital. EvolutionaryScale published its ESMFold paper in March 2023 and its ESM3 paper in January 2025, one of the key achievements being the engineering of a novel fluorescent protein. In November 2025, the team at EvolutionaryScale was acquired by the Chan Zuckerberg Initiative for an undisclosed amount.
Isomorphic Labs: Isomorphic Labs was founded in 2021 as a DeepMind spinout led by Demis Hassabis. Hassabis cofounded DeepMind in 2010, where AlphaFold was developed. Alphafold was one of the first models to predict protein structure from sequence. The key technology of Isomorphic Labs is proprietary models built on top of AlphaFold and AlphaGenome. In January 2024, Isomorphic Labs announced two major pharma partnerships with Eli Lilly and Novartis. In March 2025, Isomorphic Labs raised its first external funding round, for $600 million, led by Thrive Capital at an undisclosed valuation. As of April 2026, this is the only financing Isomorphic Labs has raised. In January 2026, it announced a partnership with Johnson & Johnson. In secondary markets, Isomorphic Lab’s valuation was estimated at $3 billion as of April 2026.
Xaira Therapeutics: Xaira Therapeutics was founded in 2023 and emerged from stealth with $1 billion in funding in April 2024 at an undisclosed valuation. David Baker, who was awarded the Nobel Prize in Chemistry in 2024 for his work in protein folding, was one of the company’s five co-founders. As of April 2026, this is the only financing Xaira Therapeutics has raised. In August 2023, the Baker Lab published a paper on RFdiffusion, a generative model for protein design. In November 2025, the group published a generative antibody design model (RFantibody) using a fine-tuned RFdiffusion network. The training code for RFantibody has been exclusively licensed to Xaira Therapeutics from the University of Washington. In secondary markets, the valuation of Xaira Therapeutics was estimated to be $2.7 billion as of April 2026.
Antibody Design
Absci: Absci is a publicly traded antibody discovery company founded by Sean McClain (CEO) in 2011. In January 2025, it announced Origin-1, a diffusion-based structure generation model that was fine-tuned from Boltz-1. Absci differentiates itself from Chai Discovery and Nabla Bio by going after the most difficult epitopes with its “zero prior epitope strategy.” The company announced a strategic $20 million investment from AMD in January 2025 to deploy its GPUs to power the AI efforts of Absci. The company has a market capitalization of $560 million as of April 2026.
Big Hat Biosciences: Big Hat Biosciences was founded in 2019 by Mark DePristo and Peyton Greenside. In 2022, Big Hat announced a $75 million Series B at an undisclosed valuation led by Section 32. Big Hat Biosciences is known for its Milliner platform, a suite of machine learning technologies integrated with a high-speed wet lab, to design and engineer therapeutic antibodies with superior functionality. In April 2025, Big Hat Biosciences announced a partnership with Eli Lilly for two antibody therapeutic programs. As of April 2026, Big Hat Bioscience has raised a total of $99.3 million.
Nabla Bio: Nabla Bio was founded in May 2020 by Surge Biswas (CEO), Frances Anastassacos, and George Church. In May 2024, it announced its $26 million Series A at an undisclosed valuation. As part of this round, Nabla Bio also announced collaborations with AstraZeneca, Bristol Myers Squibb, and Takeda. In November 2024, Nabla Bio announced its AI protein design system, JAM. In November 2025, Nabla Bio announced its second-generation antibody design system, JAM-2. The platform has achieved some of the highest hit rates (39% VHH-Fc) through large-scale validation (923+ designs). In October 2025, Nabla Bio announced the signing of its second research partnership with Takeda. The deal involved upfront and research cost payments in double-digit millions, with Nabla Bio being eligible to receive success-based payments that may exceed $1 billion. As of April 2026, Nabla Bio has raised a total of $37.4 million.
OmniAb: OmniAb is a publicly traded antibody discovery company. In November 2022, the company was spun out of Ligand Pharmaceuticals as an independent company. Its core technology is its Biological Intelligence (BI) platform, which is used to engineer the immune systems of transgenic animals to create optimized antibody candidates. The company markets multiple transgenic animal models, OmniRat, OmniChicken, and OmniMouse, each genetically engineered to produce antibodies with human sequences. OmniAb further offers OmniFlic and OmniClic, which are designed for bispecific antibody discovery. As of January 2026, the company has 104 active partners and 399 active discovery programs. OmniAb has a market capitalization of $210 million as of April 2026.
Business Model
Chai Discovery operates a hybrid business model, which involves both platform licensing and research partnerships.
Platform Licensing
The company primarily generates revenues through licensing access to its platform to biopharmaceutical companies and smaller biotech companies.
Chai-1 is open-access under an Apache-2.0 license, which gives free access for academic and commercial use. The focus of Chai-1 is to build adoption and credibility within the scientific community.
For Chai-2, customers pay an annual access fee for Chai’s AI molecule designing software.
This two-pronged approach builds scientific credibility through open-source contributions while capturing value from commercial deployments. Academic validation through open sourcing is crucial for biopharma adoption of the more premium models that Chai Discovery offers.
Research Partnerships
Chai announced a biopharma partnership with Eli Lilly in January 2026 to accelerate Eli Lilly’s internal pipeline development. Part of this deal involves training exclusive models on Eli Lilly’s proprietary data and developing new tools for Eli Lilly. Typical commercial agreements of these closed services models involve a royalty of 0.5% to 5% for the discovery contribution made by the company.
Traction
Academic Adoption
In September 2024, Chai-1 was publicly released and open-sourced. As of April 2026, Chai Discovery has accumulated over 1.9K GitHub stars and over 400 academic citations, demonstrating adoption across the structural biology community. The model is being used in leading research institutions, including the David Baker lab at the University of Washington, where the researchers reported that Chai-1 “proved remarkably effective at identifying the most active designs" for enzyme engineering applications.
Pharmaceutical Industry
In June 2025, Chai Discovery revealed the antibody design capabilities of Chai-2 and received hundreds of early access requests from pharmaceutical and biotech companies seeking to validate the platform. In November 2025, Chai Discovery published data on the results of using Chai-2 on difficult-to-drug proteins like GPCRs. Chai Discovery reported 50% hit rates with Chai-2, with its experimentally determined structures of Chai-2 designs closely matching the computational predictions.
In January 2026, Chai Discovery announced its first major biopharmaceutical partnership with Eli Lilly, which, as of April 2026, is the largest biopharma company by market capitalization. Under this collaboration, Eli Lilly pays a mid-eight-figure annual access fee to deploy Chai Discovery’s antibody design tools across its therapeutic portfolio. As part of this deal, Eli Lilly internally validated Chai Discovery’s models before signing, and Chai Discovery is training exclusive models on Eli Lilly’s proprietary internal datasets.
Multiple discovery teams in other pharmaceutical companies have confirmed active deployment of Chai Discovery’s tools in their own programs. According to a scientist at Johnson & Johnson, Chai Discovery is being used in parallel with AlphaFold for kinase inhibitor screening, biomarker discovery, and small molecule design.
Government Collaborations
In June 2025, Chai Discovery was selected as a strategic industry collaborator for the UK’s OpenBind initiative alongside Isomorphic Labs and Genentech (Roche). This initiative aims to generate the world’s largest open dataset of drug-protein interactions used for generative model training.
Valuation
In December 2025, Chai Discovery announced a $130 million Series B co-led by Oak HC/FT and General Catalyst at a $1.3 billion valuation, bringing the total funding as of April 2026 to $231 million. This round followed Chai Discovery’s Series A raise in August 2025, where it announced a $70 million Series A led by Menlo Ventures from its Anthology Fund, valuing the business at $550 million.
Chai Discovery has attracted investment from prominent individual investors in the tech industry, including Sarah Guo (Founder of Conviction), Greg Brockman (Co-founder of OpenAI), Blake Byers (Founder of New Limit), Lachy Groom (Founder of Physical Intelligence), Fred Ehrsam (Co-founder of Coinbase), Julia Hartz and Kevin Hartz (Founders of Eventbrite), Will Gaybrick (President, Technology and Business at Stripe), and Martin Chavez (Partner at Sixth Street).
Key Opportunities
Displacing High Throughput Screening
The antibody therapeutics market is made up of over 150 FDA-approved antibodies representing decades of traditional discovery methods. These conventional methods utilize high-throughput screening (HTS) methodologies, which require the investment of both time and capital. A typical HTS screening campaign costs between $500K to $2 million to generate viable antibody candidates. Further, these methods are based on libraries that have inherent biases.
Chai Discovery’s zero-shot antibody design achieved a breakthrough 20% experimental hit rate on 52 diverse targets, which represents a 200x improvement over previous computational methods, which achieved success rates in the range of 0.1%.
A traditional HTS program would require teams of five to ten people working for one to three months per target. With Chai Discovery, this process would take weeks by designing and testing generated antibody sequences. This speed advantage would translate into cost reductions for biopharma companies that spend millions of dollars on HTS, a market valued at over $20 billion.
Further, HTS is limited by the size and composition of molecule libraries, while Chai Discovery is limited by the dataset of antibody sequences that its models were trained on. This enables Chai Discovery to discover “best-in-class” molecules against known epitopes. Owning best-in-class assets is important for sustaining revenue. For example, Lipitor (atorvastatin) was the fifth statin to enter the market, but its better clinical outcomes resulted in being deemed best-in-class, allowing it to become the best-selling statin.
Expanding Asset Portfolios
Traditional discovery methods limit companies to testing a limited set of therapeutic hypotheses due to resource constraints. Chai Discovery is able to shift the discovery burden from laboratory capacity to computational resources. According to a former director at Absci, traditionally, if a company had two therapeutic targets and wanted to design a molecule that hits both, this would require a 30- to 50-person team.
This changes how therapeutic portfolios can be managed at biopharma companies. Instead of being forced to pursue therapeutic hypotheses that a company would be highly confident in, Chai Discovery enables companies to “brute force” therapeutic hypotheses that were deemed to be too risky. Meier describes this in an August 2025 interview, “Sometimes people are like, how did you decide to try this idea versus this other idea? And we’re often just trying tons of them in the lab because it’s just easy enough to do it right now.”
Expanding Platform Capabilities Beyond Antibodies
The technical infrastructure Chai Discovery has built is modality agnostic and is limited by therapeutic hypotheses. Many adjacent markets can be targeted by Chai Discovery. Peptide therapeutics, popularized by the success of GLP-1 and biohackers, can be targeted by Chai Discovery’s platform. Chai-2 achieved a 68% success rate on mini-protein designs (50 to 100 amino acids), which are slightly larger than peptides (2 to 50 amino acids). This expansion path is outlined by Meier, “We’ve shown with mini-proteins that we can get 68% success rates with picomolar affinities. There’s no reason that with other classes of molecules, those success rates can’t be that high as well.”
Other therapeutic modalities are encompassed by this as well. RNA therapeutics constituted a market exceeding $6 billion in 2023, and RNA structure modelling is clinically important for this therapeutic modality. Therapeutics like mRNA, siRNA, microRNA, IncRNA (long non-coding RNA), and riboswitches are all emerging as promising therapeutic modalities that would all benefit from structure modeling. That being said, the feasibility of RNA structure modeling is greatly limited by the amount of publicly available RNA structures.
Key Risks
Competitive Convergence
The de novo antibody design market has become increasingly crowded. This includes open-source models like RFantibody from the David Baker lab, and Boltz-2, which was developed at MIT. Further, there are closed-source competitors such as JAM-2 from Nabla Bio and Origin-1 from Absci.
According to a former Absci Director, the pace of change continues to accelerate: “Every six months there will be a new model claiming it has superiority in silico than the existing models.” Hence, any technical edge a company in this space has gets rapidly competed away.
Most companies rely on similar datasets in model training: the 200K protein structures in the Protein Data Bank (PDB) and the 10K antibody structures in the Structural Antibody Database (SAbDab). According to the former Absci Director: “There’s no moat you get from the training set. The ability to generalize from the training set or generate new sequences only now comes from your ML strategy and only comes from the compute you use.” This means Chai Discovery’s competitive advantage relies on algorithmic innovation and computational resources, which can be replicated by well-funded competitors.
The current market remains fragmented. While Chai Discovery focuses on hit generation for standard antibody discovery, competitors are carving out differentiated positions. Isomorphic Labs specializes in lead optimization with over $3 billion in platform partnerships signed, Nabla Bio targets difficult epitopes with demonstrated superiority on membrane proteins, and Absci focuses on a “zero prior epitope” strategy for novel targets.
Further, competitors are developing similar technologies. In February 2026, Isomorphic Labs announced IsoDDE, which demonstrated 3x the accuracy of Chai-1 on protein-ligand structure prediction. The model also outperformed AlphaFold 3 by 2.3x and Boltz-2 by 19.8x in the high-fidelity regime (DockQ > 0.8) on a test set of novel antibody-antigens.
Market Reality of a Platform Business Model
Chai Discovery has a platform-first business model that faces significant headwinds from historical biotech market dynamics, where the platform business model has consistently struggled to capture economic value compared to its asset-focused peers. For example, the TYK2 inhibitor developed by Nimbus Therapeutics was sold to Takeda for $4 billion in 2023. Schrödinger, a computational drug discovery platform, was the key computational platform used in the discovery, but it only captured 3.7% of the exit value ($147.3 million). As of April 2026, Schrödinger maintains a market capitalization of approximately $870 million, a quarter of the value created by the assets discovered using its platform.
This issue with value capture is structural to the platform model. A typical partnership arrangement with biopharma likely involves an industry-standard royalty rate of 0.5-5% of product net sales for molecules discovered using Chai Discovery’s platform. If the biopharma partner develops a blockbuster antibody, a drug with at least $1 billion in annual sales, Chai Discovery would receive between $5-50 million per year in royalties. However, if Chai Discovery develops the same asset internally and licenses it to the same biopharma partner at standard biotech terms (5-20% in royalties and milestone payments), it would capture more value.
The AI drug discovery companies that were founded before Chai Discovery in 2024 reflect this point. Recursion Pharmaceuticals had originally positioned itself as a platform discovery company. However, the company progressively moved towards asset development, focusing on advancing its MEK2 inhibitor through Phase II. Generate:Biomedicines, another platform competitor, has similarly developed its own clinical pipeline, with the company filing to go public to advance its assets through Phase III clinical trials.
Chai Discovery is particularly under pressure because of its $1.3 billion valuation achieved at its Series B, which is based on the value of its platform rather than its assets. If the platform business model cannot generate sufficient returns to justify this valuation, Chai Discovery may be forced to pivot towards asset development. However, this pivot is not easy. It requires additional capital to pay for clinical development costs ($100-200 million per asset) and the expansion of headcount to cover clinical, regulatory, and manufacturing.
Services Trap
Chai Discovery’s current partnership strategy is exemplified by the Eli Lilly collaboration announced in January 2026. The Eli Lilly deal explicitly includes developing “a purpose-built AI model, exclusively for use by Eli Lilly, trained on large-scale proprietary Eli Lilly data and tailored to Eli Lilly’s discovery workflows.” While this customization of services may justify premium pricing, with the deal reported to have an annual access fee in the “mid-eight figures”, it may also take away engineering resources that go into improving the core Chai Discovery platform.
As Chai Discovery continues to sign new partnerships with more major pharmaceutical companies, if these deals require the development of custom models, the company would have to support separate engineering teams to maintain separate codebases while simultaneously advancing the core platform. If Chai Discovery continues to accept partner customization, it will gradually transform into a contract research organization with limited scalability.
Proprietary Data Gap
Both Chai Discovery and its competitors are fundamentally constrained by public datasets: the 200K protein structures in the Protein Data Bank (PDB) and the 10K antibody structures in the Structural Antibody Database (SAbDab). Hence, every competitor starts from the same starting point of baseline data.
The critical missing data types are largely not found in these public databases. For example, for antibody engineering, important things to consider are immunogenicity, manufacturability, and formulation. Although most of this data is collected by biopharma companies, these companies are not incentivized to share this data with companies like Chai Discovery, which would give another biopharma competitor a competitive edge. According to a former director at Absci, there are fewer than 1K data points in public repositories for things like immunogenicity, yet properly training a robust model requires over 10K data points.
Several companies are addressing this challenge by investing resources into obtaining proprietary data through two strategies.
Internal wet lab capabilities: Competitors like Absci and Generate:Biomedicines have built substantial internal experimental capabilities to generate proprietary training data. The challenge is the cost of doing this, as purchasing DNA at scale is expensive. According to the former Absci director, this is the biggest pain point: “I really think if you were going to fix a system in a wet lab, the biggest thing to fix is the DNA purchasing. It's really expensive.”
Pharmaceutical data partnerships: Biopharma partners have decades of internal drug discovery data that can be used to train models. Many are beginning to recognize the value of this data. For example, in March 2025, Openfold, an open-source research consortium, announced a partnership with Johnson & Johnson and AbbVie to combine its data in a federated manner.
Chai Discovery’s current approach appears to rely primarily on the data of its pharmaceutical partners. In January 2026, Chai Discovery announced that it would train proprietary models for Eli Lilly using their internal data. As part of this arrangement, Chai Discovery will have access to experimental data from Eli Lilly that will be used to train better models.
Further, for Chai Discovery, it opted to forego significant internal wet lab capabilities, making it reliant on external data. Meier describes this in an interview, “Almost every AI bio company before us has had some kind of very tight lab integration with what they are doing. And it is almost too tight.” As of April 2026, Chai Discovery’s job postings don’t reflect a plan for growing an internal wet lab team.
Decade-Long Clinical Validation Gap
None of the antibodies Chai Discovery has produced has been clinically developed. Just because an antibody can bind doesn’t mean that it will not have any off-target effects. This creates a validation gap that potentially lasts 10 to 15 years before definitive proof of the clinical utility of Chai Discovery’s tools. Further, although molecules may perform well in vitro, in vivo validation introduces another layer of attrition. According to a former Johnson & Johnson scientist, “In vivo setting does not reflect 100% what we have seen in vitro, like in the lab. Once you go into the clinic or the preclinical level, that’s again out of 10 molecules, only two or three molecules work.”
The primary failure mode of assets happens in clinical development due to immunogenicity and off-target effects. Generative models may be able to circumvent these failures by having them learn the features that drive immunogenicity or developability. However, the available public data that models can learn these features from is extremely scarce. If early clinical results from Chai-2-designed antibodies reveal unexpected toxicity, immunogenicity, or efficacy failures, the perception of these computationally designed biologics would suffer.
Further, the central challenge for the AI drug discovery industry is that no AI-discovered molecule has completed clinical development. The value of a computation platform is tied to the assets that come out of it, and an asset’s value is only realized once it becomes fully clinically approved. For Chai Discovery, realizing clinical milestones is the only path to demonstrating progress in its platform.
Summary
Chai Discovery is a venture-backed AI biotech company that builds generative models for de novo antibody design. Its flagship Chai-2 model achieved 20% experimental hit rates across 52 de-novo designed targets, a 200x improvement over prior computational methods. Founded in March 2024 by a team with roots in OpenAI and Absci, the company has gained traction both with its open-source Chai-1 structure prediction model and its pharmaceutical partnership with Eli Lilly.
However, Chai Discovery faces material risks. Competition in the generative modeling space is heating up, with rivals like Isomorphic Labs, Xaira Therapeutics, Nabla Bio, and Absci closing technical gaps using similar public training data. All while the platform services business model remains historically unproven for the industry.